For background, please read the post on the
K=48 analysis and links therein.
As I mentioned in the previous posts, my technique depends on the use of MDS to reduce the dimensionality of genomic data from 177,000 SNPs or so to a few dozen dimensions capturing most of the variance.
Subsequently,
mclust a state of the art clustering algorithm is applied on the MDS representation: this iterates between choices of K, the number of clusters, trying clusters of different shape, volume, and orientation, and chooses the
optimal clustering, maximizing the Bayes Information Criterion. In simpler terms, it finds as much detail as possible in the data but penalizes too ornate models and avoids finding "ghost" clusters that are not really supported.
These are clusters derived from data of unlabeled individuals. The only human input into the process is the number of MDS dimensions to retain.
In my previous K=48 analysis, I retained 30 dimensions, but I also noted that this is not really optimal. Choosing more, or less, dimensions might lead to even better resolution (higher K).
More dimensions = more possible ways to distinguish between individuals, but also, possibly, more noise, as individuals might not be "clustered" in them.
Fewer dimensions = less possible ways to distinguish between individuals, but also, possibly, less noise from the uninformative higher dimensions.
Thus, the question arises: how many dimensions to retain?
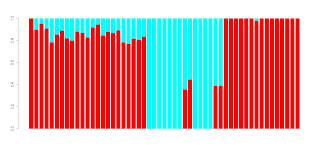
Here is a plot of the optimal number of clusters inferred, depending on how many dimensions I chose to retain:
As you can see, when a few dimensions are retained, relatively few clusters are inferred, while as the number of dimensions goes beyond a certain point, the number of clusters starts to decrease again, as more noise is added (*)
The number of clusters peaks (see figure) at 16 and 22 dimensions retained; both of these produce 56 different clusters in the optimal solution.
Here are the results for Dodecad Project members (up to DOD236) with K=56 and 16 dimensions retained. In comparison to the previous K=48 analysis, we are now able to:
- Split CEU White Utahns (#1) from French (#15)
- Split CEU White Utahns (#1) from continental Germanics (#14)
- Split French (#15) from Spaniards (#2)
- Split Armenians (#7) from Turks (#19)
- Split Slavs (#23) from Balts (#26)
- Split Cypriots (#30) from Sephardic Jews (#21)
The most astonishing finding is, however, at least for me,
the emergence of a cluster (#16) comprised in the great majority by people from Greece and Southern Italy, with very few individuals from elsewhere. Notice that #16 has absolutely no representation in the reference populations, which lack South Italians and Greeks.
Once again, I urge participants to help themselves and others by leaving a comment in the
ancestry thread.
(*) The change is not, however, smooth. The more general problem is to choose which dimensions to retain, rather than choosing how many of the first ones to retain. The first few dimensions of MDS capture a decreasing portion of variance, but the data are not guaranteed to be "split" in them. However, this is a much harder problem, as we have to figure out (i) how many dimensions to retain, and (ii) which ones. Even if we fix (i), by choosing to retain, e.g., 10 dimensions, we still have to choose which 10: this is close to half a billion different combinations of which 10 to choose from the total of 38 possible candidate ones.